This introduction first discusses how homology is inferred from significant similarity, and how those inferences can be confirmed, and then considers strategies that connect homology to more accurate functional prediction. The inference of functional similarity from homology is more difficult, both because functional similarity is more difficult to quantify, and because the relationship between homology (structure) and function is complex. While similarity searching is an effective and reliable strategy for identifying homologs – sequences that share a common evolutionary ancestor – most similarity searches seek to answer a much more challenging question: "Is there a related sequence with a similar function?". The units in this chapter present practical strategies for identifying homologous sequences in DNA and protein databases (units 3.3, 3.4, 3.5, 3.9, 3.10) once homologs have been found, more accurate alignments can be built from multiple sequence alignments (unit 3.7), which can also form the basis for more sensitive searches, phenotype prediction, and evolutionary analysis. Similarity searching is effective and reliable because sequences that share significant similarity can be inferred to be homologous they share a common ancestor. (1997) units 3.3 and 3.4), PSI-BLAST ( Altschul et al., 1997), SSEARCH ( Smith and Waterman (1981) Pearson (1991), unit 3.10), FASTA ( Pearson and Lipman (1988) unit 3.9) and the HMMER3 ( Johnson et al., 2010) programs produce accurate statistical estimates, ensuring protein sequences that share significant similarity also have similar structures. Widely used similarity searching programs, like BLAST ( Altschul et al. Modern protein sequence databases are very comprehensive, so that more than 80% of metagenomic sequence samples typically share significant similarity with proteins in sequence databases. Sequence similarity searching to identify homologous sequences is one of the first, and most informative, steps in any analysis of newly determined sequences. Staphylococcus aureus coagulase restriction fragment length polymorphism sequence-based phylogenetic analysis.AN INTRODUCTION TO IDENTIFYING HOMOLOGOUS SEQUENCES The coa genotypes and their restriction patterns observed in the present study are novel, not published earlier. Two coa patterns were observed in mastitic milk indicating multiple origins of infection, with 595 bp coa genotype being predominant in mastitic milk. aureus coagulase typeshave a site-specific predilection. The study, being localized to only one farm, yielded different RFLP patterns as observed from different sampling sites, which indicates that different S. While as the most distant sequences with the value of 0.483 were found between S.
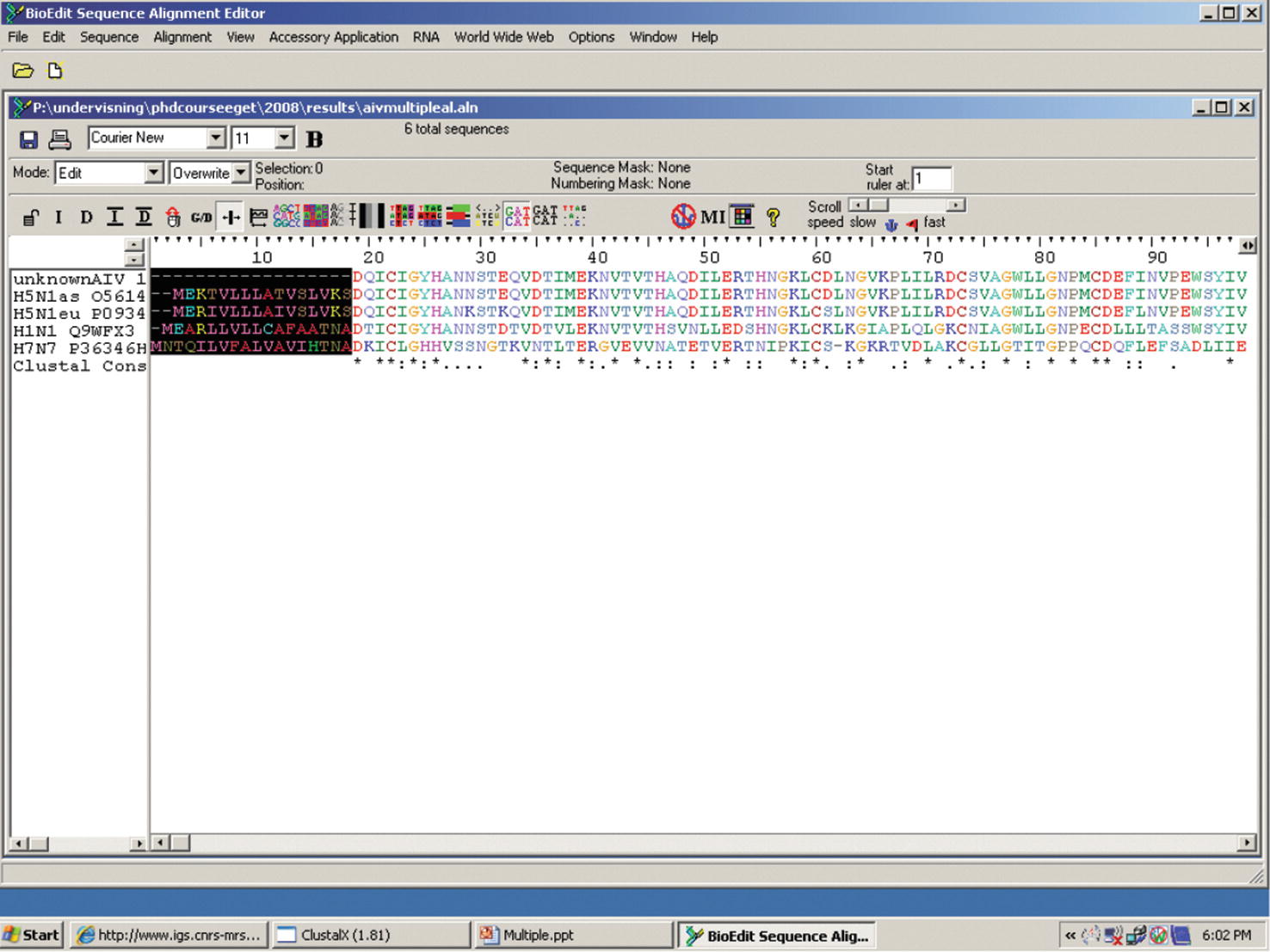
aureus 595 (mastitic milk), and thus, they are most closely related. The most identical sequences with the value of 0.810 were found between isolate S. On sequencing, four different sequences having unique restriction patterns were obtained. On RFLP using both restriction enzymes, four different restriction patterns P1, P2, P3, and P4 were observed.
#BIOEDIT SEQUENCE IDENTITY MATRIX SKIN#
514 bp (2 isolates) and 757 bp (4 isolates) coa genotypes were observed from nasal cavity and pus from skin wounds, respectively. Two coa genotypes, 595 bp (15 isolates) and 802 bp (4 isolates), were observed in mastitic milk. Four different genotypes of coa gene, i.e., 514 bp, 595 bp, 757 bp, and 802 bp were obtained. aureus isolates, 25 (64.10%) isolates carried coa gene. aureus were confirmed by targeting nuc gene using PCR. Of 192 different samples,39 (20.31%) isolates of S. These sequences were aligned for maximum homology using the Bioedit softwareandsimilarity in the sequences was inferred with the help of sequence identity matrix. One isolate from each restriction pattern was sequenced. Different coa genotypes observed were subjected to RFLP using restriction enzymes Hae111 and Alu1, to obtain the different restriction patterns. aureus isolates were subjected to coagulase ( coa) gene PCR. The presumptive isolates were confirmed by nuc gene-based polymerase chain reaction (PCR). This study was conducted to study the coagulase gene-based genetic diversity of Staphylococcus aureus, isolated from different samples of cattle using restriction fragment length polymorphism (RFLP) and their sequence-based phylogenetic analysis.Ī total of 192 different samples from mastitic milk, nasal cavity, and pus from skin wounds of cattle from Military Dairy Farm, Jammu, India, were screened for the presence of S.
